A single-cell multiomics roadmap of zebrafish spermatogenesis
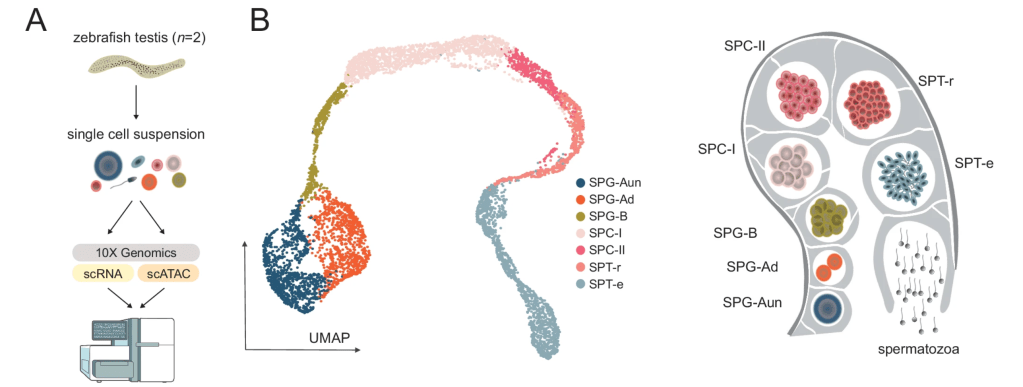
We are excited to share our latest work, now published in Molecular Systems Biology, led by Ana María Burgos-Ruiz. In this study, we present a comprehensive single-cell multiomics atlas of zebrafish spermatogenesis, integrating scRNA-seq, scATAC-seq, and base-resolution DNA methylation (WGBS) across defined germ cell populations. This resource provides a high-resolution view of transcriptional and epigenomic dynamics underlying male germline development in an anamniote vertebrate. We identify the major germ cell states and key regulatory drivers that shape their progression, alongside stage-specific chromatin accessibility and DNA methylation changes. Notably, we observe widespread chromatin compaction in spermatids, coupled with a subset of loci that remain accessible—highlighting candidate regions that may carry regulatory information across generations. Beyond advancing our understanding of zebrafish germline biology, this work establishes a framework for comparative analyses of spermatogenesis across vertebrates and offers insights into potential mechanisms of epigenetic inheritance.